United Kingdom (EN)
Select your region or country.
Pushing the limits of bioluminescence microscopy
Bioluminescent microscopy’s unique advantages are pushing researchers to challenge it against conventional fluorescent methods. When detecting and imaging live cells and animals, the use of bioluminescence reporters has shown to be a promising method, especially in neuroscience research.
Dr. Michael Krieg, from ICFO, Institut de Ciencies Fotòniques, Castelldefels, Spain, and his collaborators demonstrate volumetric imaging of fast cellular dynamics by exposing the advantages and pushing the limits of bioluminescence microscopy on Caenorhabditis elegans and other biological model organisms.

Caenorhabditis elegans is a tiny (1 mm in length) nematode which has been used as a model for the last 60 years due to its stereotypic body plan, transparent skin, and well-characterized nervous system. Copyright © 2022, Luis Felipe Morales-Curiel, Michael Krieg et al.
The advantages and challenges of bioluminescence
In bioluminescence microscopy, the signal is generated by a chemical reaction of an enzyme (luciferase) with its substrate (luciferin). The general approach and the information content of bioluminescence microscopy are similar to that of fluorescence microscopy. However, one significant advantage is that bioluminescence does not use excitation light. This increases the specificity of the signal since endogenous autofluorescence will not create any background signal, potentially hiding the specific signal under investigation.
Another advantage is the reduced stress of the sample, since the intense excitation light – as used in fluorescence microscopy, might induce some unwanted, potentially toxic effects in the sample, as can be seen in Application Example 1.

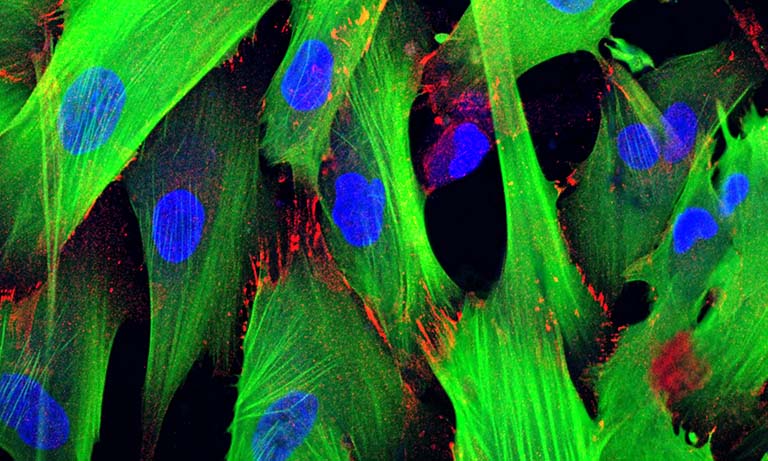
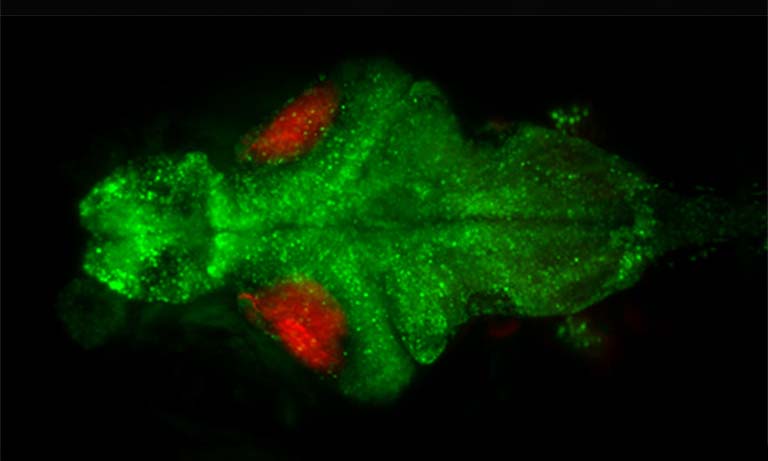
Application Example 1 : Fluorescence vs bioluminescence on C. elegans after induced stress1;
The cellular stress response of C. elegans was measured using a mNeonGreen-NanoLantern fused to DAF-16, which acts as a stress reporter. Without any stressors, DAF-16 is located in the cytoplasm, however, under stress, DAF-16 is mostly found in the nucleus. The experiment was performed using fluorescence microscopy of the mNeonGreen (panel b) as well as bioluminescence using the mNeonGreen-Nanolantern fusion (panel c). The ratio of signal from the nucleus to the cytoplasm is shown in panel d. Following a heat shock, the stress response is detected both with fluorescence as well as bioluminescence, but the effect is more prominent with bioluminescence detection, likely due to the absence of non-specific autofluorescence. Interestingly the control experiment (shown in the insets in b and c) indicates a stress response for fluorescence imaging, most likely due to the exposed excitation radiation.
Fluorescent markers also tend to bleach over time, limiting long-time observation of live samples. Those effects are efficiently mitigated using bioluminescence microscopy, which obtains the energy required for the signal generation from chemical processes rather than the photophysical process involved in fluorescence.
Nonetheless, a major drawback of bioluminescence is its rather low signal intensity, resulting in long exposure times and low SNR images in microscopy, limiting its applicability in dynamic processes, which are of big interest when observing living specimens.
The experimental setting

Dr. Michael Krieg and his Graduate student, Luis Felipe Morales-Curiel using the highly light-efficient microscope “LowLiteScope”
In his experiment, Dr. Michael Krieg and his group optimized every aspect of the experimental pipeline for bioluminescence. Using extremely photon-starved samples to challenge these limitations, they aimed to provide high SNR images at high temporal resolution. To achieve this, he used Nanolanterns for the dyes, which are brighter and more varied in their spectral properties than the classically used luciferases. Since a standard widefield microscope did not provide any signal, they built a highly light-efficient microscope termed “LowLiteScope”.
Additionally, as the resulting images were still difficult to inspect due to the low signal available, content aware image restoration (CARE) was applied to improve these, which provided clear, high contrast images at millisecond exposure times.
Imaging with quantitative CMOS technology
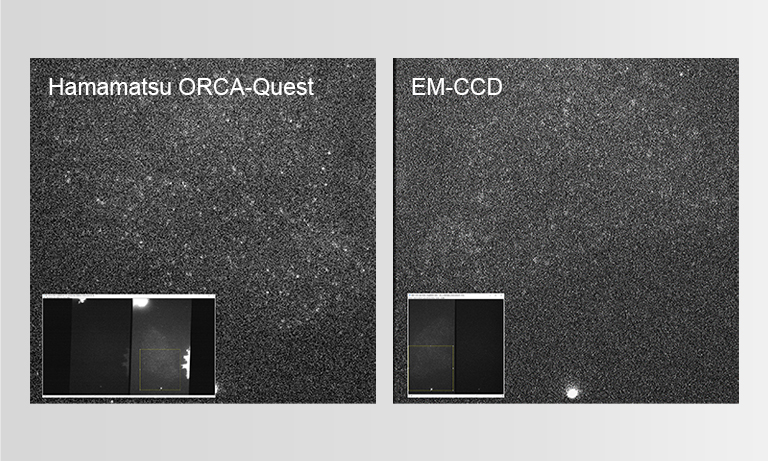
To ensure the most efficient detection, the quantitative CMOS (qCMOS) camera ORCA-Quest was used to offer the best images from the available photons. Its extremely low readout noise of 0.27 e-, alongside its high quantum efficiency, provides the optimal signal-to-noise ratio down to almost the single photon level.
Apart from the raw sensitivity, another advantage of the ORCA-Quest in the presented bioluminescence setting lies within its geometrical design: in combination with the optics of the LowLiteScope, the pixel pitch of 4.6 μm provides a perfect tradeoff between light collection efficiency and spatial resolution.
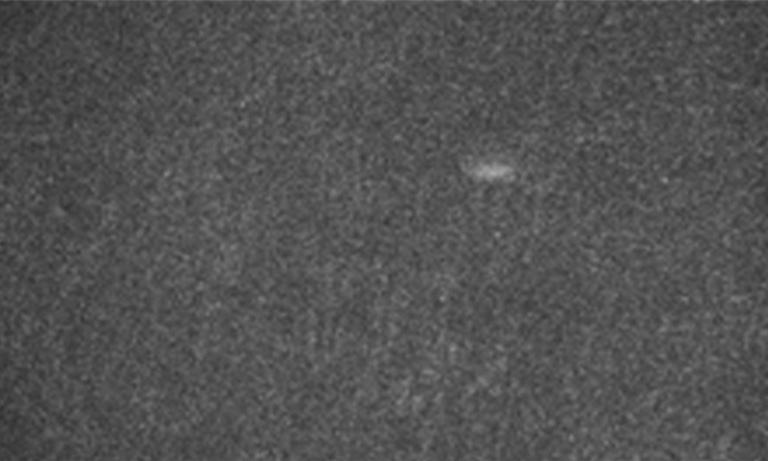
Since the ORCA-Quest delivers a pixel count of 9.4 MP, no sacrifices in regard to the field of view needed to be made. The pixel pitch and pixel count also benefitted another evolution of the method: in order to increase the speed of 3D acquisitions, lightfield microscopy was also included. The small pixel sizes and the high pixel count of the ORCA-Quest allowed the projecting of the 3D lightfield onto the two dimensional camera sensor, enabling rapid 3D imaging of the whole C. elegans using bioluminescence. Classical lightfield imaging requires a complicated computational reconstruction of the 3D object which might take up to 30 minutes. To speed up this process, a neural network was employed, reducing the time for 3D reconstruction down to 100 ms. Using this scheme, satisfactory results can be achieved with exposure time down to 5 s. To further reduce the minimum exposure time in order to image sample dynamics in three dimensions, CARE was applied before the reconstruction. This reduced the achievable exposure times further up to a factor of 20 while keeping acceptable exposure times. An example of bioluminescence AI reconstructed lightfield microscopy at 5 volumes per second is shown in Application Example 2 below.

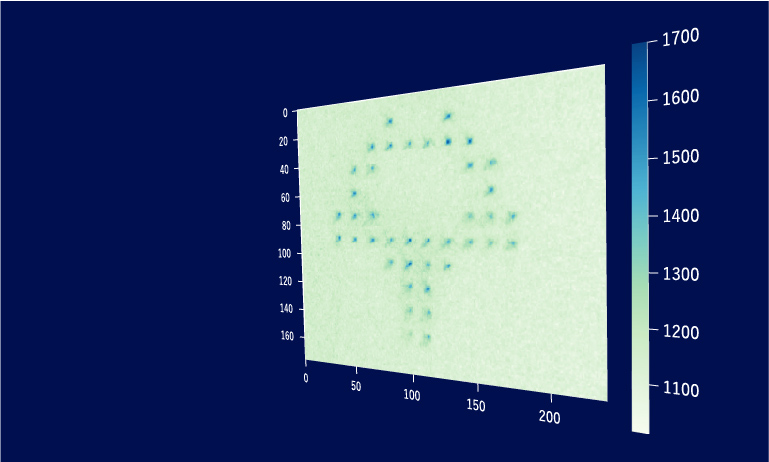
Application Example 2 : Raw lightfield images of a bioluminescent calcium reporter in muscles of a freely moving C. elegans
Results and future applications
Luis Felipe Morales-Curiel and Dr. Michael Krieg have now established bioluminescence microscopy as a versatile addition to the biologist toolbox. Bioluminescence offers the benefits over fluorescence of lower stress to the samples as well as higher specificity. The common drawbacks of bioluminescence techniques, typically being low signal yield and resulting long exposure times were overcome by optimizing the whole imaging pipeline. This included using the brightest bioluminescence dyes available, recording the data with instrumentation optimized for low light sensitivity, and finally processing the data with modern machine learning based approaches.
In the Krieg lab, the bioluminescence framework will be used to further study mechanobiology and neuroscience of C. elegans, other model organisms but also stem cell-derived organoids. In a recent study2, this framework has been applied to visualize the performance of calcium-triggered photon emitters for establishing a photon-based synaptic communication pathway.
About Dr. Michael Krieg

Dr. Michael Krieg was a Postdoctoral Research Fellow in the Department of Molecular and Cellular Physiology at Stanford University. In the lab of Dr. Miriam Goodman, Michael investigated basic mechanotransduction pathways in neurons of the model organism, C. elegans. Michael earned his Ph.D. in Developmental Biology/Biophysics from the Technical University Dresden. Currently, Dr. Krieg is the Group Leader of the Neurophotonics and Mechanical Systems Biology Research Group at the Institute of Photonic Sciences, ICFO, Spain.
Sources
- https://www.nature.com/articles/s42003-022-04292-x
- https://www.nature.com/articles/s41592-023-01836-9
- Read the full research paper here: https://rdcu.be/c9Re5
Product used in this application & case study
The C15550-20UP is the world's first camera to incorporate the qCMOS image sensor. The camera achieves the ultimate in quantitative imaging.
Other Application & Case study
- Confirmation
-
It looks like you're in the . If this is not your location, please select the correct region or country below.
You're headed to Hamamatsu Photonics website for GB (English). If you want to view an other country's site, the optimized information will be provided by selecting options below.
In order to use this website comfortably, we use cookies. For cookie details please see our cookie policy.
- Cookie Policy
-
This website or its third-party tools use cookies, which are necessary to its functioning and required to achieve the purposes illustrated in this cookie policy. By closing the cookie warning banner, scrolling the page, clicking a link or continuing to browse otherwise, you agree to the use of cookies.
Hamamatsu uses cookies in order to enhance your experience on our website and ensure that our website functions.
You can visit this page at any time to learn more about cookies, get the most up to date information on how we use cookies and manage your cookie settings. We will not use cookies for any purpose other than the ones stated, but please note that we reserve the right to update our cookies.
1. What are cookies?
For modern websites to work according to visitor’s expectations, they need to collect certain basic information about visitors. To do this, a site will create small text files which are placed on visitor’s devices (computer or mobile) - these files are known as cookies when you access a website. Cookies are used in order to make websites function and work efficiently. Cookies are uniquely assigned to each visitor and can only be read by a web server in the domain that issued the cookie to the visitor. Cookies cannot be used to run programs or deliver viruses to a visitor’s device.
Cookies do various jobs which make the visitor’s experience of the internet much smoother and more interactive. For instance, cookies are used to remember the visitor’s preferences on sites they visit often, to remember language preference and to help navigate between pages more efficiently. Much, though not all, of the data collected is anonymous, though some of it is designed to detect browsing patterns and approximate geographical location to improve the visitor experience.
Certain type of cookies may require the data subject’s consent before storing them on the computer.
2. What are the different types of cookies?
This website uses two types of cookies:
- First party cookies. For our website, the first party cookies are controlled and maintained by Hamamatsu. No other parties have access to these cookies.
- Third party cookies. These cookies are implemented by organizations outside Hamamatsu. We do not have access to the data in these cookies, but we use these cookies to improve the overall website experience.
3. How do we use cookies?
This website uses cookies for following purposes:
- Certain cookies are necessary for our website to function. These are strictly necessary cookies and are required to enable website access, support navigation or provide relevant content. These cookies direct you to the correct region or country, and support security and ecommerce. Strictly necessary cookies also enforce your privacy preferences. Without these strictly necessary cookies, much of our website will not function.
- Analytics cookies are used to track website usage. This data enables us to improve our website usability, performance and website administration. In our analytics cookies, we do not store any personal identifying information.
- Functionality cookies. These are used to recognize you when you return to our website. This enables us to personalize our content for you, greet you by name and remember your preferences (for example, your choice of language or region).
- These cookies record your visit to our website, the pages you have visited and the links you have followed. We will use this information to make our website and the advertising displayed on it more relevant to your interests. We may also share this information with third parties for this purpose.
Cookies help us help you. Through the use of cookies, we learn what is important to our visitors and we develop and enhance website content and functionality to support your experience. Much of our website can be accessed if cookies are disabled, however certain website functions may not work. And, we believe your current and future visits will be enhanced if cookies are enabled.
4. Which cookies do we use?
There are two ways to manage cookie preferences.
- You can set your cookie preferences on your device or in your browser.
- You can set your cookie preferences at the website level.
If you don’t want to receive cookies, you can modify your browser so that it notifies you when cookies are sent to it or you can refuse cookies altogether. You can also delete cookies that have already been set.
If you wish to restrict or block web browser cookies which are set on your device then you can do this through your browser settings; the Help function within your browser should tell you how. Alternatively, you may wish to visit www.aboutcookies.org, which contains comprehensive information on how to do this on a wide variety of desktop browsers.
5. What are Internet tags and how do we use them with cookies?
Occasionally, we may use internet tags (also known as action tags, single-pixel GIFs, clear GIFs, invisible GIFs and 1-by-1 GIFs) at this site and may deploy these tags/cookies through a third-party advertising partner or a web analytical service partner which may be located and store the respective information (including your IP-address) in a foreign country. These tags/cookies are placed on both online advertisements that bring users to this site and on different pages of this site. We use this technology to measure the visitors' responses to our sites and the effectiveness of our advertising campaigns (including how many times a page is opened and which information is consulted) as well as to evaluate your use of this website. The third-party partner or the web analytical service partner may be able to collect data about visitors to our and other sites because of these internet tags/cookies, may compose reports regarding the website’s activity for us and may provide further services which are related to the use of the website and the internet. They may provide such information to other parties if there is a legal requirement that they do so, or if they hire the other parties to process information on their behalf.
If you would like more information about web tags and cookies associated with on-line advertising or to opt-out of third-party collection of this information, please visit the Network Advertising Initiative website http://www.networkadvertising.org.
6. Analytics and Advertisement Cookies
We use third-party cookies (such as Google Analytics) to track visitors on our website, to get reports about how visitors use the website and to inform, optimize and serve ads based on someone's past visits to our website.
You may opt-out of Google Analytics cookies by the websites provided by Google:
https://tools.google.com/dlpage/gaoptout?hl=en
As provided in this Privacy Policy (Article 5), you can learn more about opt-out cookies by the website provided by Network Advertising Initiative:
http://www.networkadvertising.org
We inform you that in such case you will not be able to wholly use all functions of our website.
Close